Topology-Aware-Bronchial-Tree-Segmentation-via-Skeleton-Based-Branch-Parsing
Topology-aware bronchial tree segmentation from CT. Airway masks are extracted using TotalSegmentator, then skeletonized with Kimimaro. Branch points are detected and parsed using rule-based anatomical constraints to produce consistent airway labeling and multi-segment outputs for 3D Slicer.
🫁 Bronchial Branch Labeler
Automatically parse an unsegmented bronchial tree into anatomically meaningful airway branches using skeleton-based topology analysis.
📌 Overview
This project provides a lightweight, non-deep-learning pipeline to convert a binary bronchial tree into structured anatomical segments (e.g., Trachea, RMB, RUL, B1–B10).
Unlike neural network approaches, this method is:
- Deterministic
- Topology-aware
- Interpretable (based on graph structure and geometry)
It is particularly useful for:
- Airway anatomy analysis
- Preprocessing for radiology AI pipelines
- Generating labeled airway maps from CT-derived segmentations
🧠 Pipeline
CT scan (.nii.gz) ↓ TotalSegmentator ↓ Extract bronchial tree (binary mask / Segment_1) ↓ Skeletonization (kimimaro) ↓ Graph-based branch tracing ↓ Anatomical classification (B1–B10) ↓ Export:
.seg.nrrd (3D Slicer compatible) .txt (segment names)
🖼️ Demo
Bronchial branch parsing
Below shows how the algorithm converts an unsegmented bronchial tree into labeled anatomical branches:
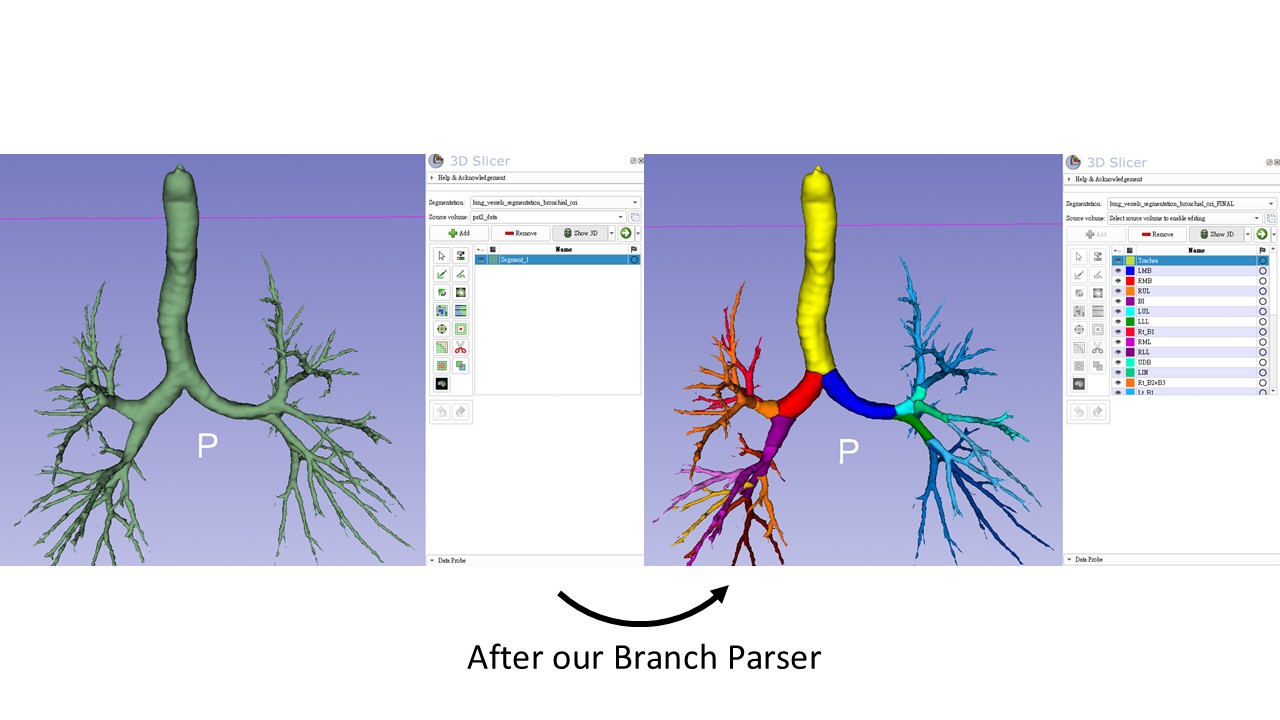
Clinical visualization benefit
The colorized airway branches significantly improve interpretability when reviewing CT in different views:
- Axial
- Coronal
- Sagittal
This allows faster identification of bronchial segments (e.g., B1–B10) during image navigation.
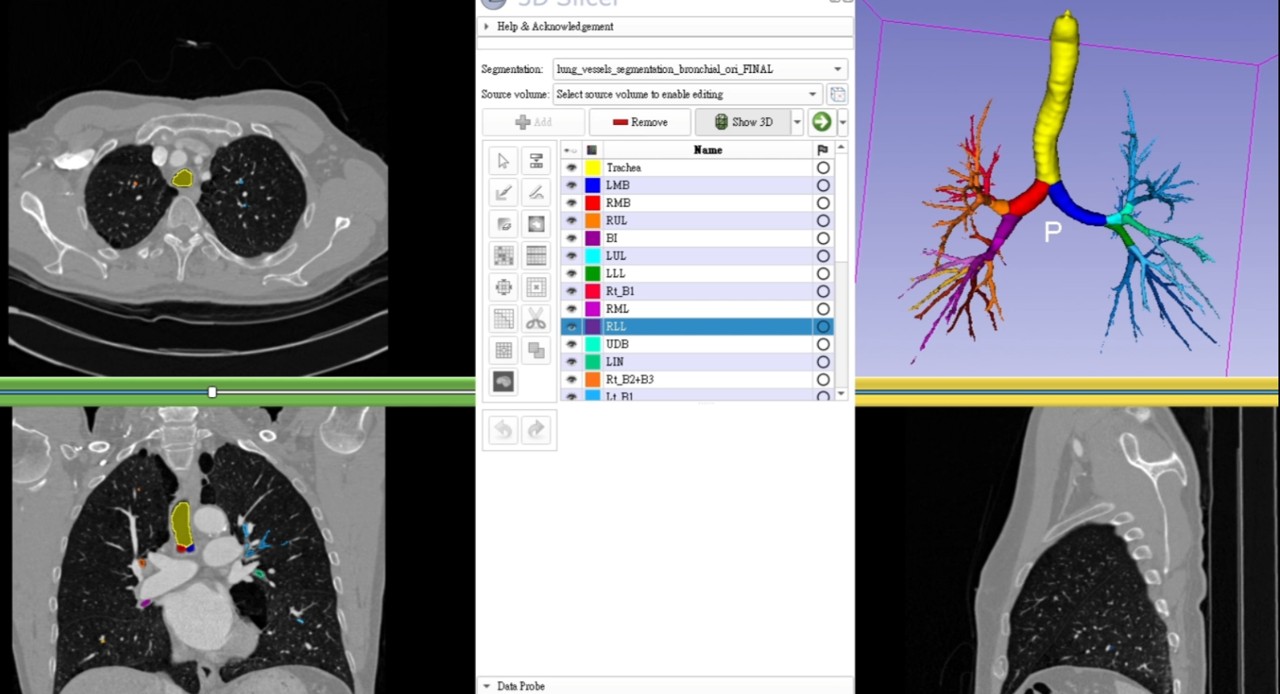
🎥 Video demonstration:
https://youtu.be/xXrjiTo91TU
⚙️ Installation
pip install numpy scipy scikit-learn nibabel pynrrd kimimaro
🚀 Usage
python bronchial_branch_labeler.py ^
--input "D:/LyNoS_dataset/Benchmark/Pat2/lung_vessels_segmentation/lung_vessels_segmentation_bronchial_ori.seg.nrrd" ^
--ct "D:/LyNoS_dataset/Benchmark/Pat2/pat2_data.nii.gz" ^
--output "D:/LyNoS_dataset/Benchmark/Pat2/lung_vessels_segmentation/bronchial_BRANCHES.seg.nrrd"
Arguments
Argument Description –input Binary bronchial tree segmentation (.seg.nrrd) –ct Original CT volume (.nii.gz) for spatial reference –output Output segmented airway file (.seg.nrrd) –ap-sign (optional) Set to -1 if anterior–posterior direction is flipped
📤 Output
- Segmentation file *.seg.nrrd Compatible with 3D Slicer Each airway branch is a separate segment layer
- Segment name file *_segment_names.txt Contains ordered anatomical labels
- Console output ```bash === FINAL SEGMENTS ===
- Trachea (bid=0)
- LMB (bid=1)
- RMB (bid=2)
- RUL (bid=3) …
- Lt_B10 (bid=38) ```
🔬 Method Highlights
Skeletonization via kimimaro Graph traversal (BFS, shortest path) PCA-based orientation estimation Branch classification using geometric heuristics Cranial direction → B1 Anterior/Posterior → B2 / B3 Spatial clustering for distal branches Topology-preserving labeling
No deep learning model is used.
⚠️ Notes
Input bronchial tree should be reasonably clean (e.g., from TotalSegmentator) Extremely noisy segmentations may affect branch detection Left B7 is not defined (consistent with anatomical convention)
📄 License
MIT License
🙌 Acknowledgement
TotalSegmentator for whole-body CT segmentation kimimaro for skeletonization
💡 Future Work
Support for airway variants Integration with radiology AI pipelines Quantitative airway analysis (diameter, angle, topology metrics)